¶ Welcome to the GlyComb help page
GlyComb is a novel glycoconjugate data repository to assign a unique identifier for each glycoconjugate dataset, including glycopeptides, glycoproteins, glycolipids, and glycosides.
In the life sciences, many public data repositories have been rapidly developed, including PRIDE and PeptideAtlas in proteomics and GlyTouCan and GlycoPOST in glycomics, to promote sharing and accumulating valuable data from researchers under the FAIR (Findable, Accessible, Interoperability, and Re-usable) data principles. GlyTouCan in glycomics, for example, can assign a unique identifier for each glycan structure. Such unique identifiers would be helpful because they would be able to link entries in heterogeneous databases. However, one of the missing pieces is the accumulation of glycoconjugate data. To assign such unique identifiers to glycoconjugate datasets, we started to develop this new data repository.
Currently, researchers can register glycopeptides and glycoproteins datasets to GlyComb by specifying an amino acid sequence or UniProt ID, glycosylation sites, and glycan information in the Excel or TSV file format. For the glycan structure input format, we adopted GlyTouCan IDs and the composition notation used in Byonic.
GlyComb is a member of the GlyCosmos project and it will collaborate with the other data repositories in the project such as GlyTouCan and GlycoPOST soon. For example, if the input glycan structures are not registered in GlyTouCan yet, GlyComb will automatically register them to GlyTouCan during the validation and accession number assignment process.
All submitted datasets will be normalized and strictly verified to confirm whether or not the identifiers have already been assigned to them previously. Furthermore, because GlyComb is built on top of Semantic Web technologies, users will be able to collect the specified glycoconjugate-associated information easily using the SPARQL query language. As the first glycoconjugate data repository, we expect that GlyComb will play a significant role in connecting glycomics data with other omics disciplines.
In this documentation, the overall data submission flow and available functionalities in GlyComb are described in detail. Instead, you can also watch the video (https://www.youtube.com/watch?v=-1-MUYW_wtk) demonstrating the usage of this repository on YouTube.
¶ Login and logout
¶ How to log in to GlyComb
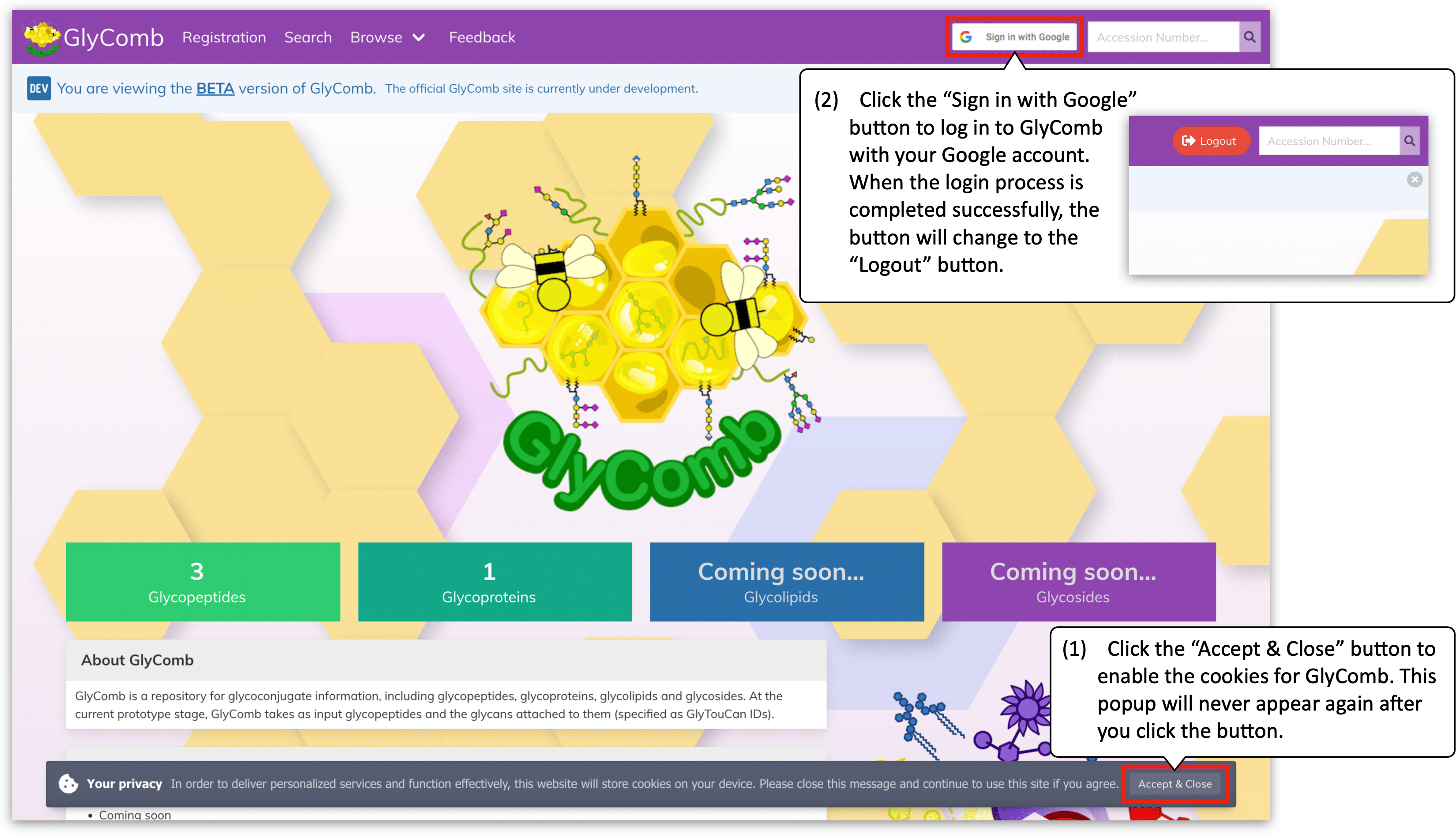
First of all, access to https://glycomb.beta.glycosmos.org. You will see the popup notifying the cookie policy at the bottom of the screen. Click the "Accept & Close" button to enable the cookies for GlyComb. This popup will never appear again after you click the button.
You can log in to GlyComb with your Google account. Please click the "Sign in with Google" button on the navigation menu then you will be redirected to the login form to Google. Fill in your email address and password to the form. When the login process completed successfully, you will be redirected to the welcome screen of GlyComb automatically and the login button on the navigation menu will change to the "Logout" button.
¶ How to log out of GlyComb
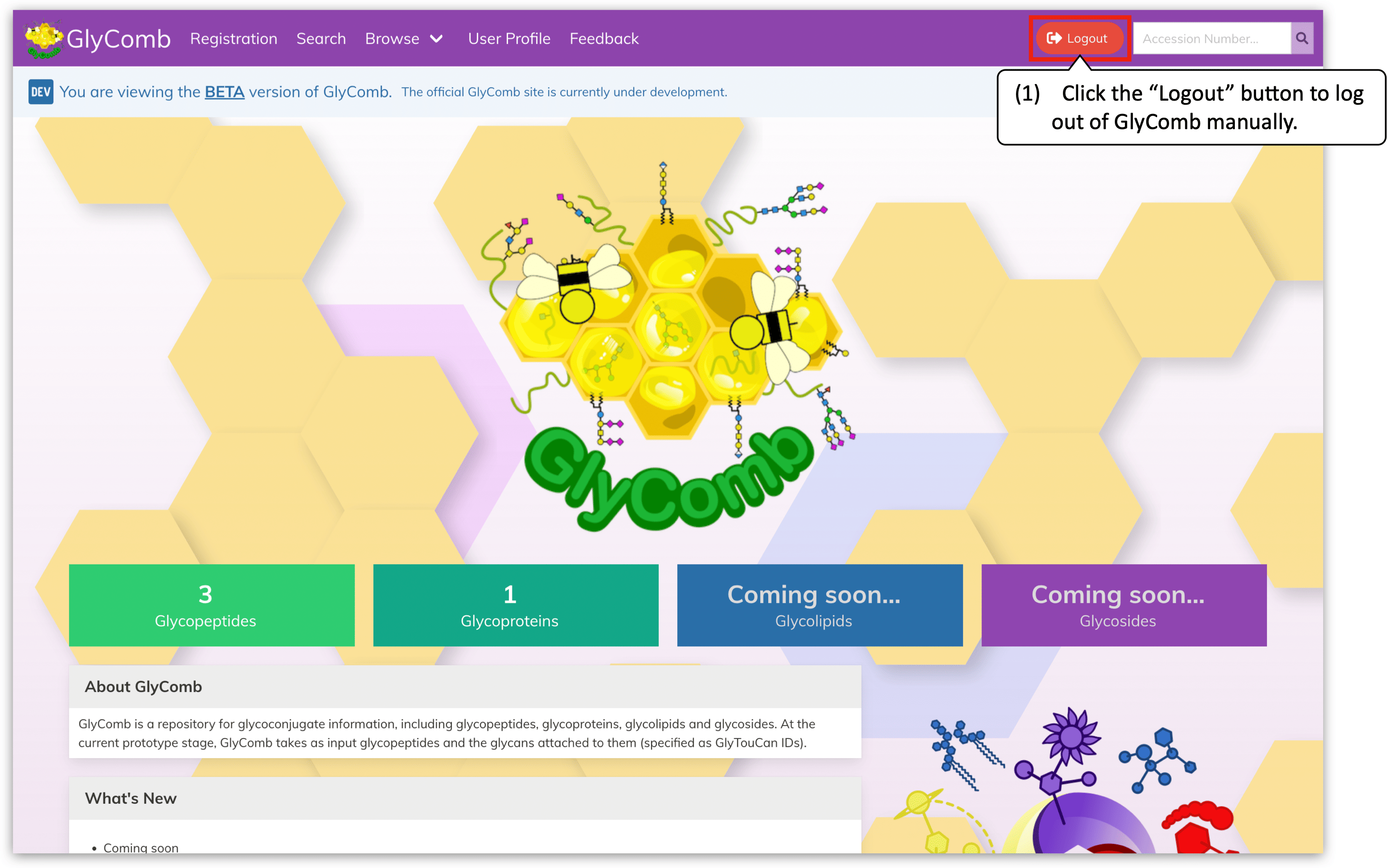
The user login session is due to expire in three hours automatically. To log out of GlyComb manually, please click the "Logout" button on the navigation menu.
¶ Browsing and searching accessions
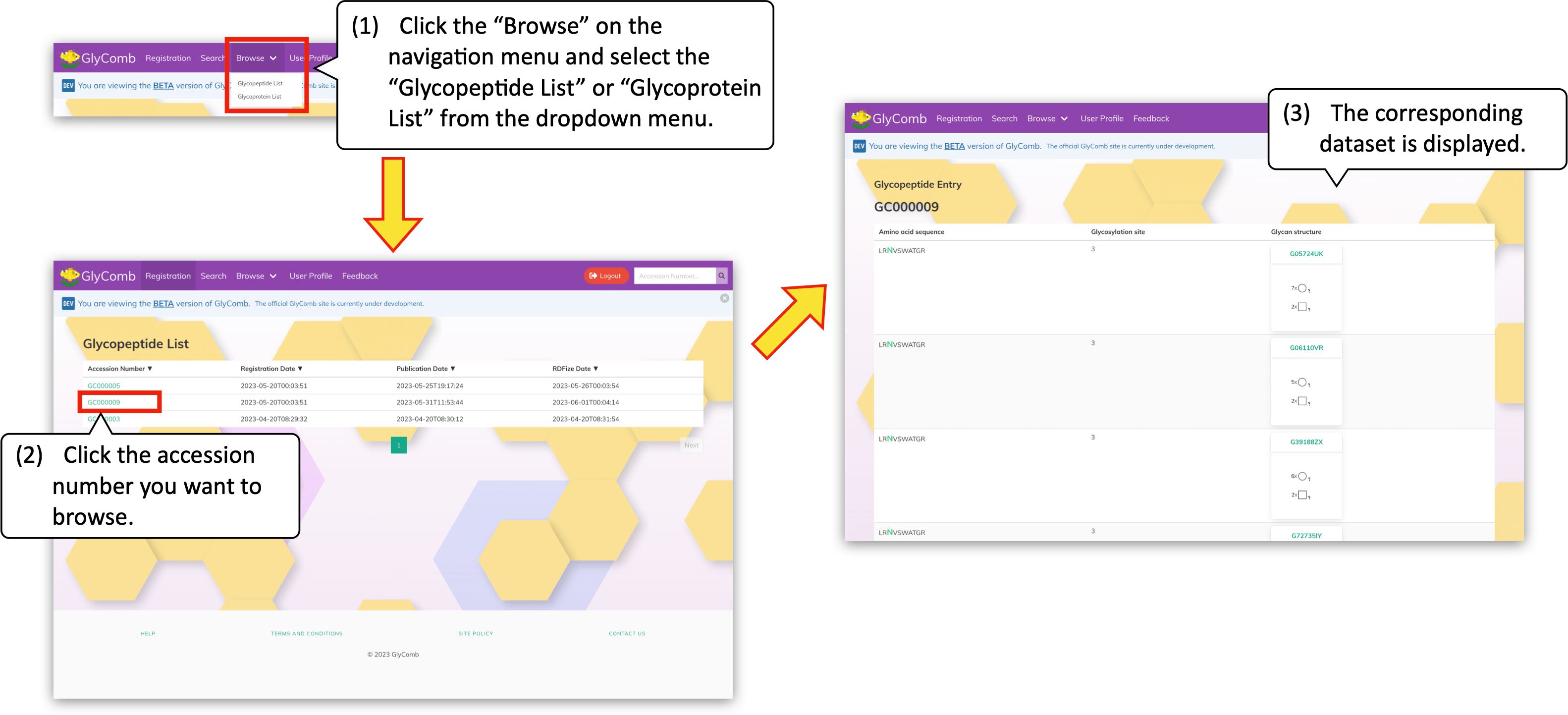
To browse the accession list, please click the "Browse" on the navigation menu. Then the dropdown menu to select whether the "glycopeptides" or the "glycoproteins" appears. By clicking an item from the menu, the accession list is displayed. You can browse the content of each accession by clicking the accession number on the list.
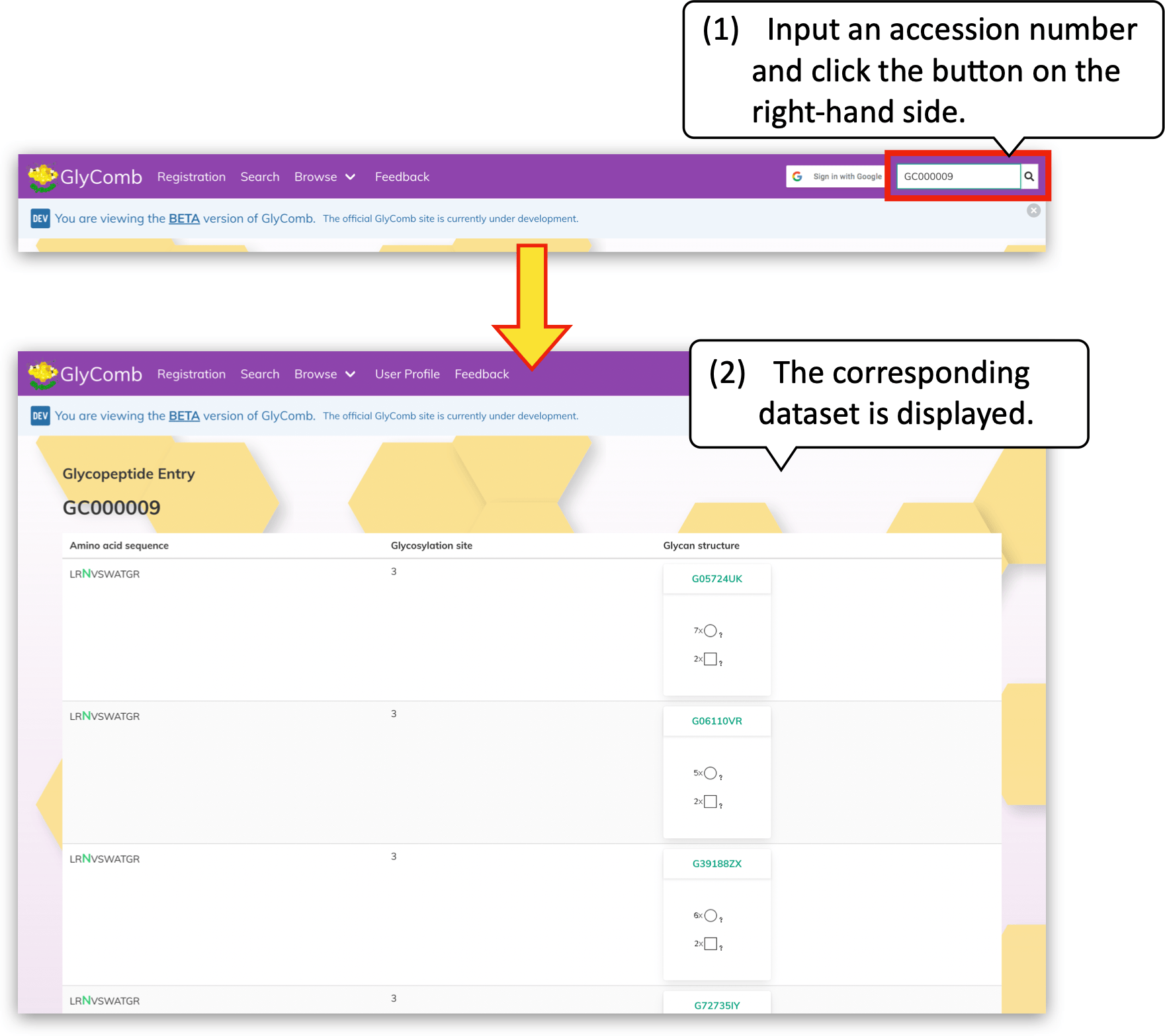
If you want to browse the dataset having a specific accession number, please enter the accession number in the text field at the top-right of the screen and click the "🔍" button instead.
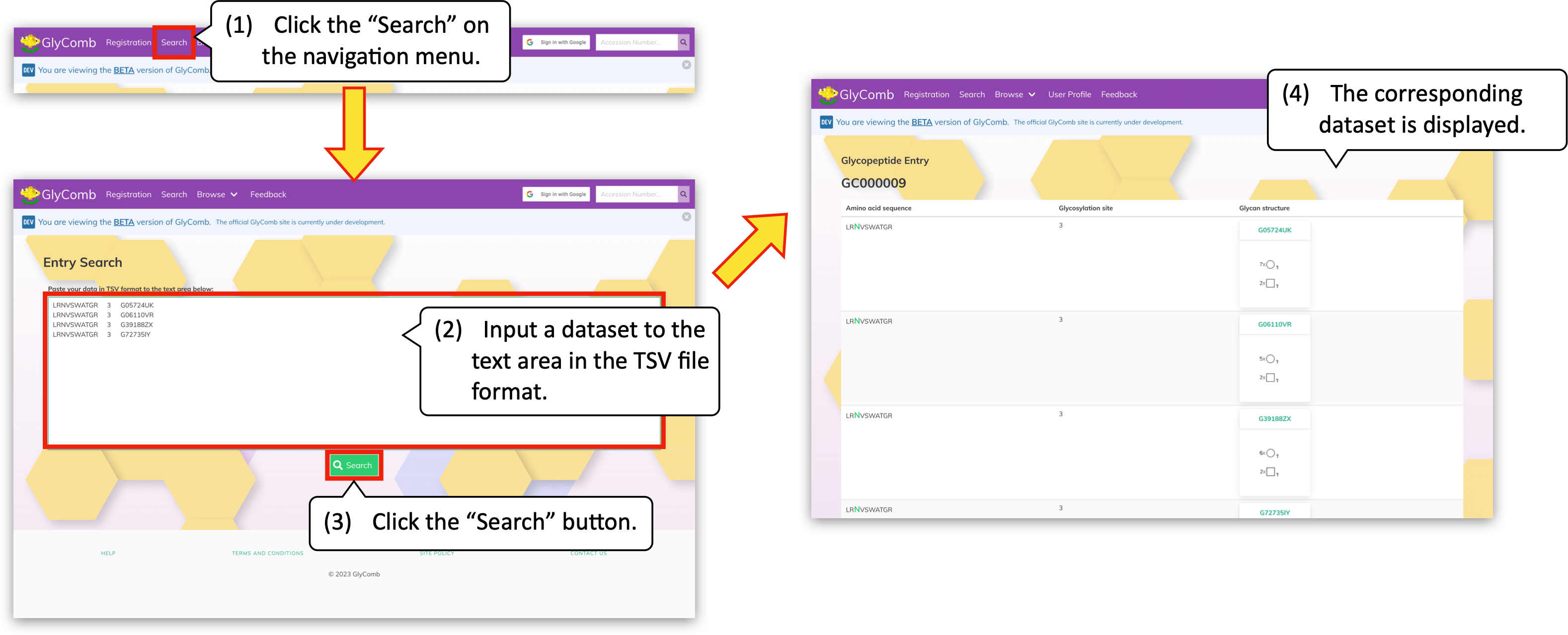
You can also search a dataset to check whether it is already registered or not. The text area to enter a dataset will be displayed by clicking the "Search" on the navigation menu. Please input a dataset in the text area in TSV file format. For more details about the format, please refer to the "Dataset submission" section below.
Then by clicking the "Search" button below, the corresponding accession will be displayed if it is already registered to GlyComb. Otherwise, an error message popup will be displayed.
¶ Dataset submissions
In this section, we describe the way to submit your glycoconjugate datasets to GlyComb. To begin with, we explain the glycoconjugate dataset format in detail. Then, the dataset submission flow and how to make the submissions available publicly are described in order.
¶ The format of the datasets
In GlyComb, a glycoconjugate dataset is composed of a substrate (e.g., a peptide sequence for the glycopeptides) and a set of the glycosylation information (i.e., the glycosylation sites and the glycan structures).
The GlyComb submission system offers two ways to submit glycoconjugate datasets: via a text area form and file uploading. From the text area form, users can submit the dataset from the clipboard in the TSV (Tab Separated Values) file format. If you prefer to submit your dataset via file upload, the following file formats are available at the time of this writing: the Excel file format (.xlsx, .xls) and the TSV file format (.tsv).
Because the formats for each glycoconjugate molecule are slightly different, thus we will describe each of them in order using the example datasets.
¶ For glycopeptide datasets
The glycopeptide datasets must be composed of three or four columns. Each of the columns contains the following data:
- The first column contains the substrate, that is to say, the peptide sequences (amino acid sequences).
- The second column contains the glycosylation site numbers. If a site number is unknown, please specify a
?character instead. - The third column contains the glycan structures. We adopted the GlyTouCan IDs and the Byonic-like composition notation to describe the glycan structures.
- The fourth column contains the UniProt IDs for supplemental information. This column is optional.
If you submit your dataset by file uploading, you can freely select the columns to read each data in the dataset file easily on the submission screen. Therefore, you don't need to worry about the order of the columns in your datasets in such cases.
The following is an example dataset that uses GlyTouCan IDs to characterize the glycan structures. This dataset is composed of three columns because it does not give any UniProt IDs.
FTFTSHTPGEHQICLHSNSTK 18 G00681RS
FTFTSHTPGEHQICLHSNSTK 18 G11617NC
FTFTSHTPGEHQICLHSNSTK 18 G19243MH
FTFTSHTPGEHQICLHSNSTK 18 G28182LP
FTFTSHTPGEHQICLHSNSTK 18 G39448FX
FTFTSHTPGEHQICLHSNSTK 18 G52182RP
FTFTSHTPGEHQICLHSNSTK 18 G54675NM
FTFTSHTPGEHQICLHSNSTK 18 G58609GW
FTFTSHTPGEHQICLHSNSTK 18 G61514UE
FTFTSHTPGEHQICLHSNSTK 18 G92463DS
The following dataset uses the Byonic-like composition notation to describe the glycan structures. This dataset is composed of four columns because it gives UniProt IDs for each line.
LRNVSWATGR 3 HexNAc(2)Hex(5) P08571
LRNVSWATGR 3 HexNAc(2)Hex(6) P08571
LRNVSWATGR 3 HexNAc(2)Hex(7) P08571
LRNVSWATGR 3 HexNAc(3)Hex(6) P08571
In addition, you can submit multiple datasets at a time by inserting a blank line for each amino acid sequence. The following is such a set of example datasets:
FTFTSHTPGEHQICLHSNSTK 18 G00681RS
FTFTSHTPGEHQICLHSNSTK 18 G11617NC
FTFTSHTPGEHQICLHSNSTK 18 G19243MH
FTFTSHTPGEHQICLHSNSTK 18 G28182LP
FTFTSHTPGEHQICLHSNSTK 18 G39448FX
FTFTSHTPGEHQICLHSNSTK 18 G52182RP
FTFTSHTPGEHQICLHSNSTK 18 G54675NM
FTFTSHTPGEHQICLHSNSTK 18 G58609GW
FTFTSHTPGEHQICLHSNSTK 18 G61514UE
FTFTSHTPGEHQICLHSNSTK 18 G92463DS
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G00681RS
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G11617NC
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G19243MH
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G28182LP
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G39448FX
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G52182RP
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G54675NM
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G58609GW
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G61514UE
QYGSEGRFTFTSHTPGEHQICLHSNSTK 25 G92463DS
SHTPGEHQICLHSNSTK 14 G00681RS
SHTPGEHQICLHSNSTK 14 G11617NC
SHTPGEHQICLHSNSTK 14 G19243MH
SHTPGEHQICLHSNSTK 14 G28182LP
SHTPGEHQICLHSNSTK 14 G39448FX
SHTPGEHQICLHSNSTK 14 G52182RP
SHTPGEHQICLHSNSTK 14 G54675NM
SHTPGEHQICLHSNSTK 14 G58609GW
SHTPGEHQICLHSNSTK 14 G61514UE
SHTPGEHQICLHSNSTK 14 G92463DS
These example datasets are in the TSV file format. The following image describes the example datasets in the Excel file format.
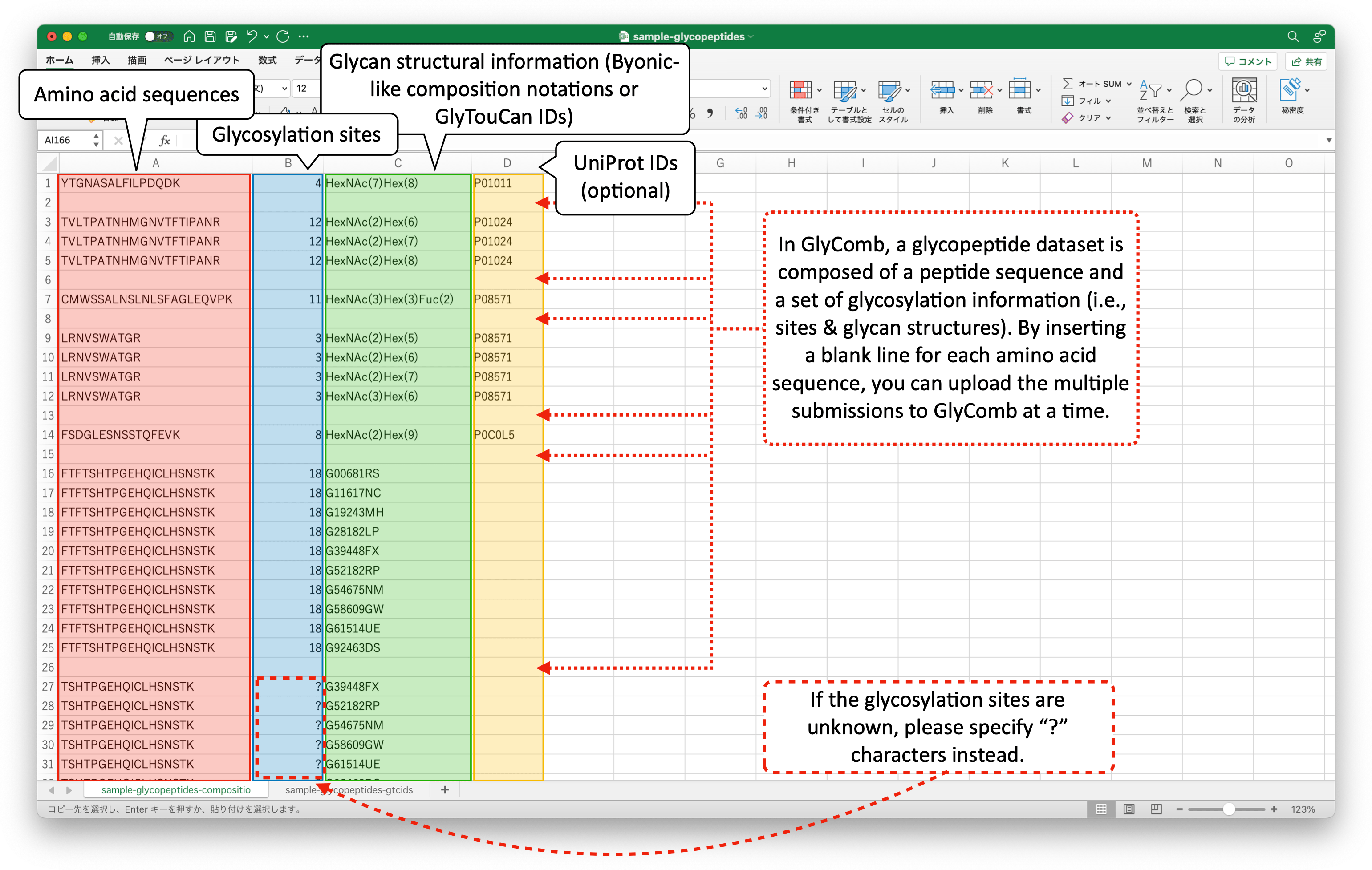

If the glycosylation site information is described as a lower-case letter in the upper-case amino acid sequence like "HIGHAnLTFEQLR" in your datasets, you can automatically extract this site information by submitting the dataset using file uploading. For more detail, please refer to the "How to submit your datasets" subsection below.
¶ For glycoprotein datasets
The glycoprotein datasets must be composed of three columns. Each of the columns contains the following data:
- The first column contains the substrate, that is to say, the UniProt IDs.
- The second column contains the glycosylation site numbers. If a site number is unknown, please specify a
?character instead. - The third column contains the glycan structures. You can specify the GlyTouCan IDs and the Byonic-like composition notation to describe the glycan structures just like the glycopeptide datasets.
If you submit your dataset by file uploading, you can freely select the columns to read each data in the dataset file easily on the submission screen. Therefore, you don't need to worry about the order of the columns in your datasets in such cases.
The following is an example dataset that uses GlyTouCan IDs to characterize the glycan structures.
Q8WWI5 135 G39188ZX
Q8WWI5 180 G06110VR
Q8WWI5 652 G49108TO
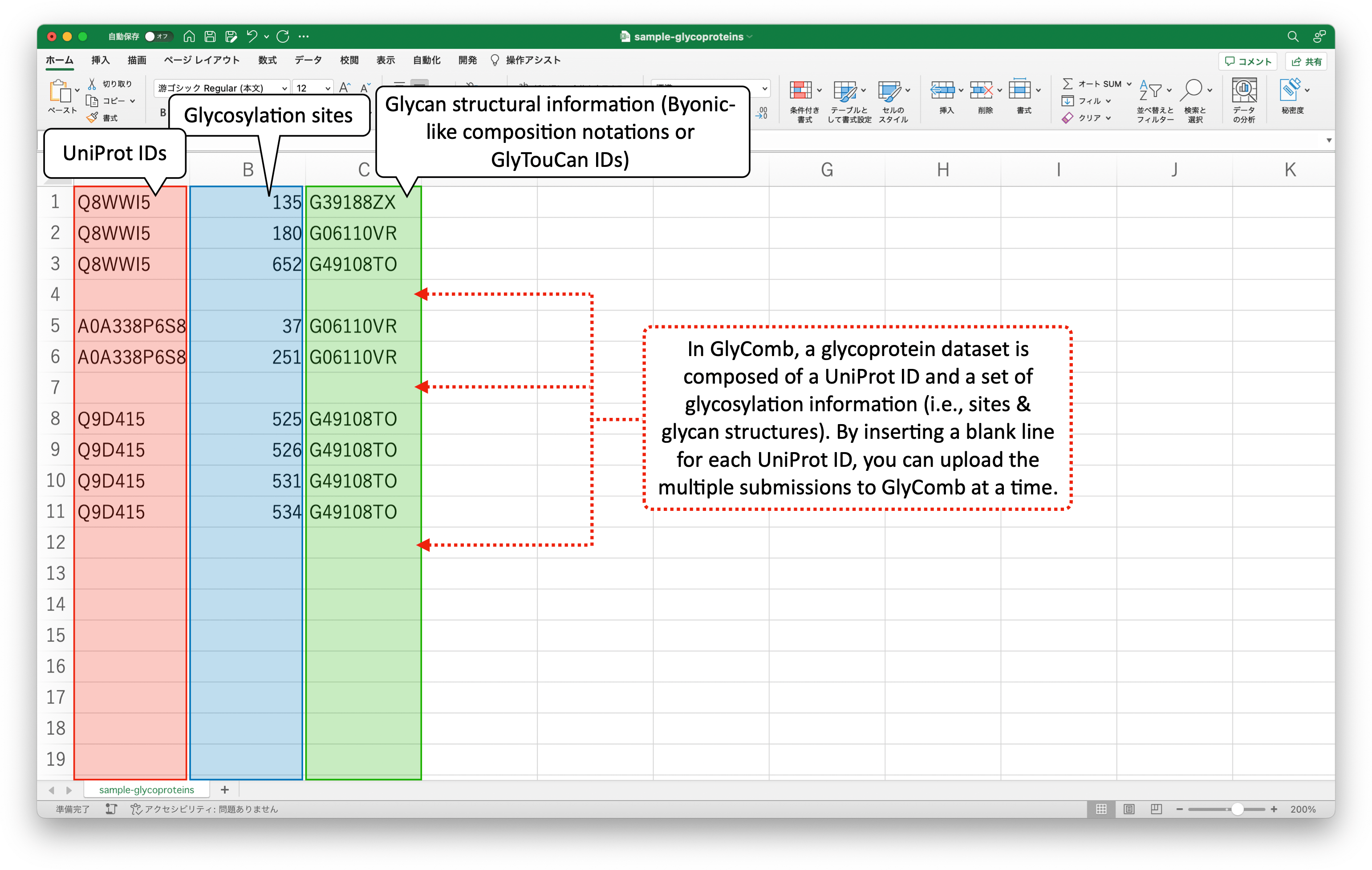
You can submit multiple datasets at a time by inserting a blank line for each UniProt ID. The following is such a set of example datasets:
Q8WWI5 135 G39188ZX
Q8WWI5 180 G06110VR
Q8WWI5 652 G49108TO
A0A338P6S8 37 G06110VR
A0A338P6S8 251 G06110VR
Q9D415 525 G49108TO
Q9D415 526 G49108TO
Q9D415 531 G49108TO
Q9D415 534 G49108TO
These example datasets are in the TSV file format. The following image describes the example datasets in the Excel file format.
¶ Byonic-like glycan composition notation
GlyComb uses slightly different notations for the substituents in the Byonic-like glycan composition notations compared to Byonic. The following table summarizes the difference between both formats. For example, HexNAc(1)Hex(2)Phospho(1) written in Byonic format is written as HexNAc(1)Hex(2)P(1) in GlyComb.
| Byonic format | Byonic-like format in GlyComb |
|---|---|
| Acetyl | Ac |
| Me- | Me |
| Phospho | P |
| Sulfo | S |
¶ How to submit your datasets
To access to the datasets registration screen, please click the "Registration" on the navigation menu. In the registration screen, select your dataset type, i.e., the "Glycopeptides" or the "Glycoproteins" from the tab menu. Then, select the "Input from clipboard" from the vertical menu if you prefer to submit your datasets via the text area form. After you enter or paste your dataset to the text area on the left-hand side of the menu, please click the "Submit" button below.
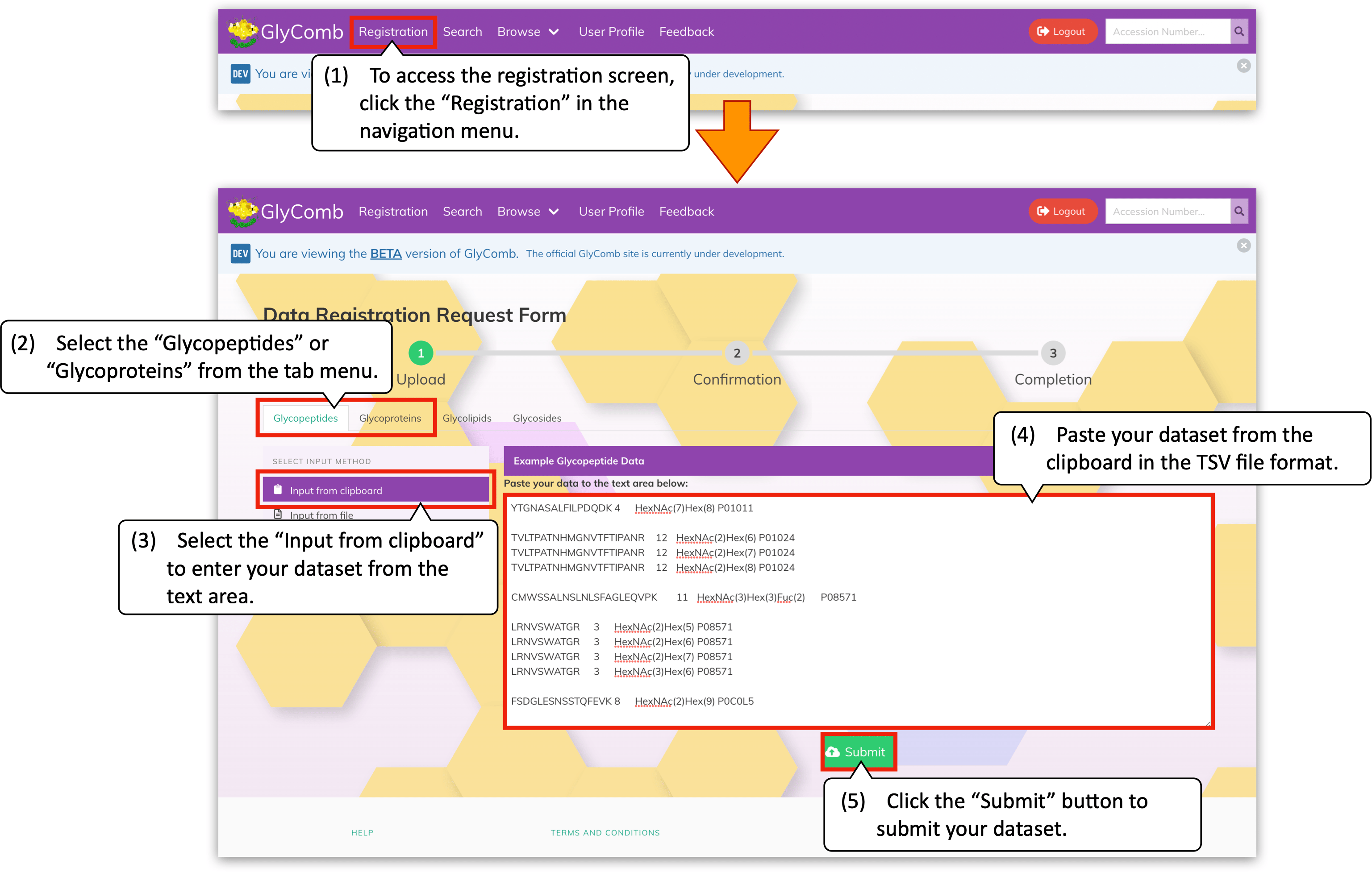
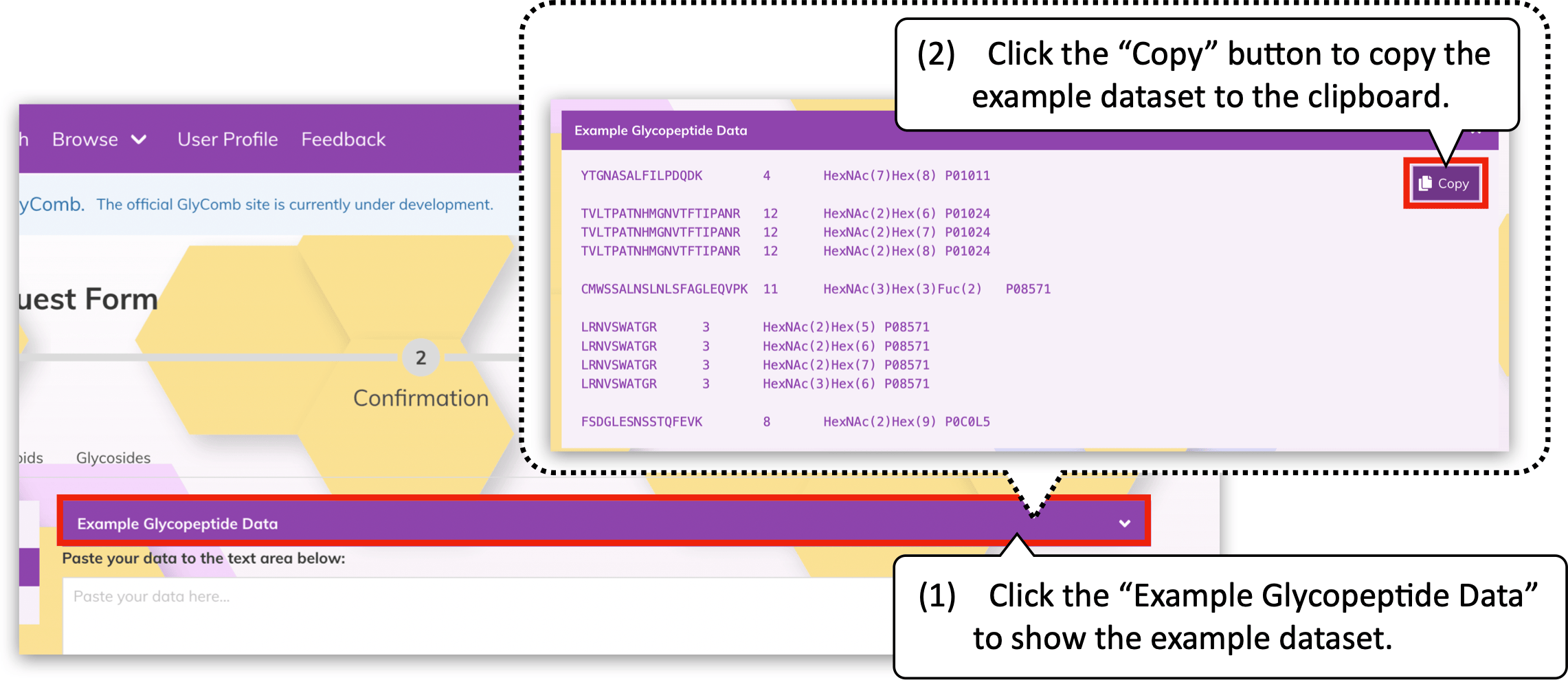
In addition, you can browse the example datasets by clicking the panel above the text area, e.g., the "Example Glycopeptide Data" for glycopeptides. When the mouse cursor hover on the example datasets, the "Copy" button will appear. You can easily copy its content to the clipboard by clicking the button.
If you prefer to submit your dataset by file uploading, please select the "Input from file" from the vertical menu in the registration screen. By clicking the "Select a file..." on the right-hand side of the menu, the file chooser dialog will appear and you can select a file containing your datasets in the Excel file format (.xlsx or .xls) or the TSV file format (.tsv).
Once you choose the dataset file, the file upload form will expand to configure more detailed dataset range, i.e., the worksheet name for the Excel files, the row number from which your dataset starts, etc. In addition, you can freely select the columns to read the dataset in this form. After you complete the dataset range configurations, please click the "Submit" button below.
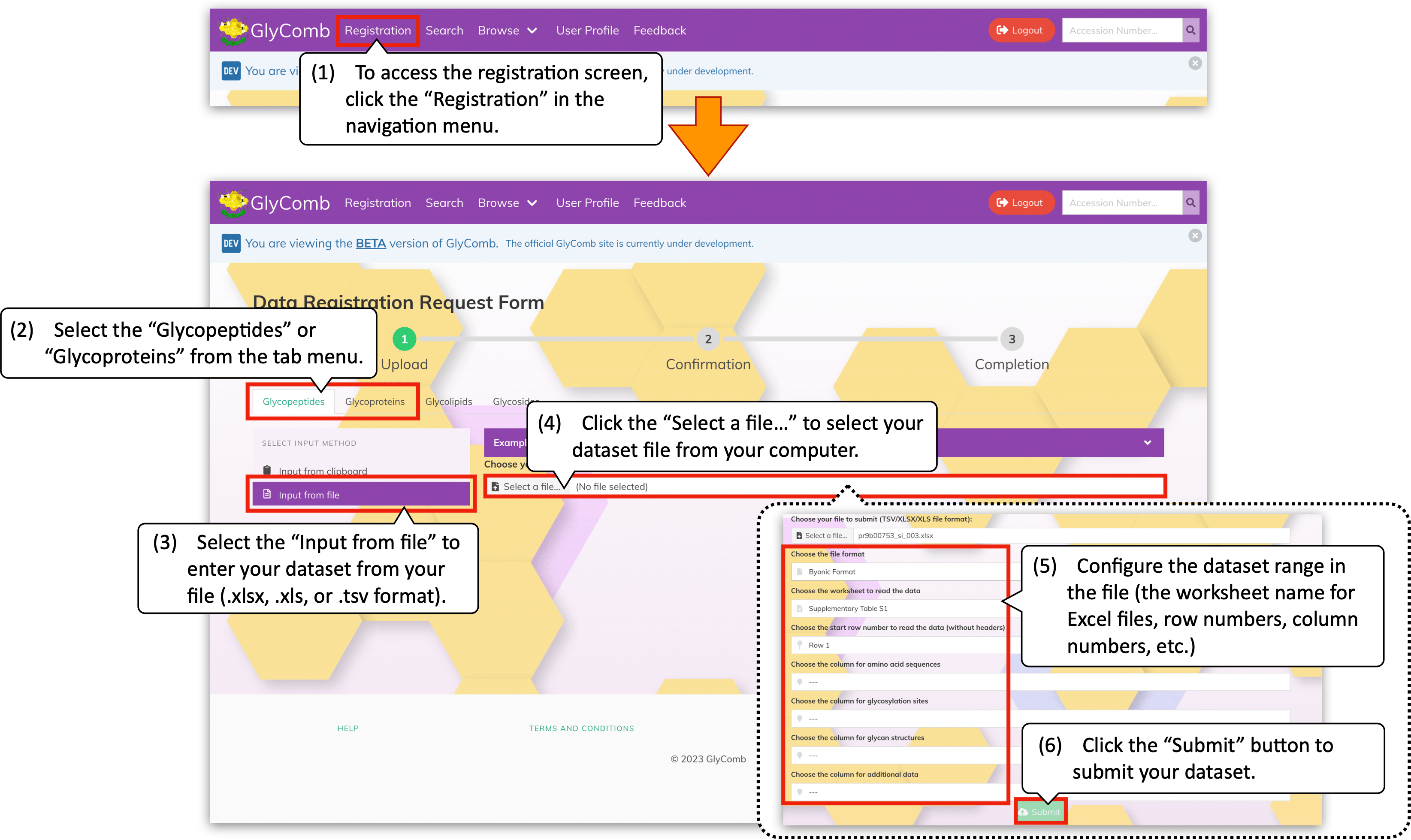
As we described in the "For glycopeptide datasets" subsubsection, you can extract the glycosylation site information embedded in the amino acid sequences like "HIGHAnLTFEQLR" automatically in this configuration form. For example, the following image describes the datasets taken from Jing et al. (2020) (https://pubs.acs.org/doi/10.1021/acs.jproteome.9b00753).
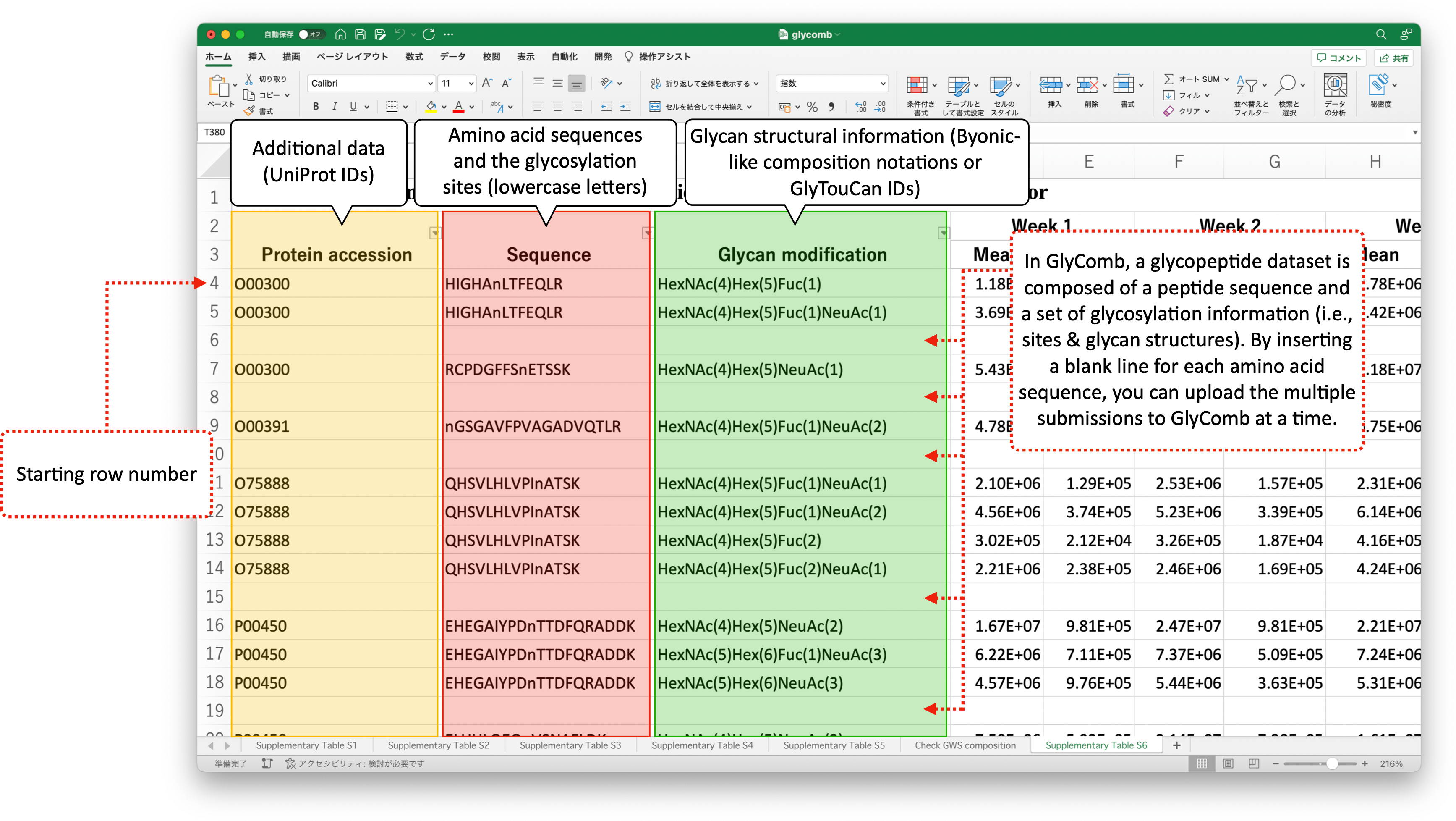
There are multiple amino acid sequences so we added a blank line for each amino acid sequence. In these datasets, the glycosylation site information is encoded as the lower-case characters in the amino acid sequences. Also, the actual datasets start from line no.4. To read such glycopeptide datasets, please configure the range like the below:
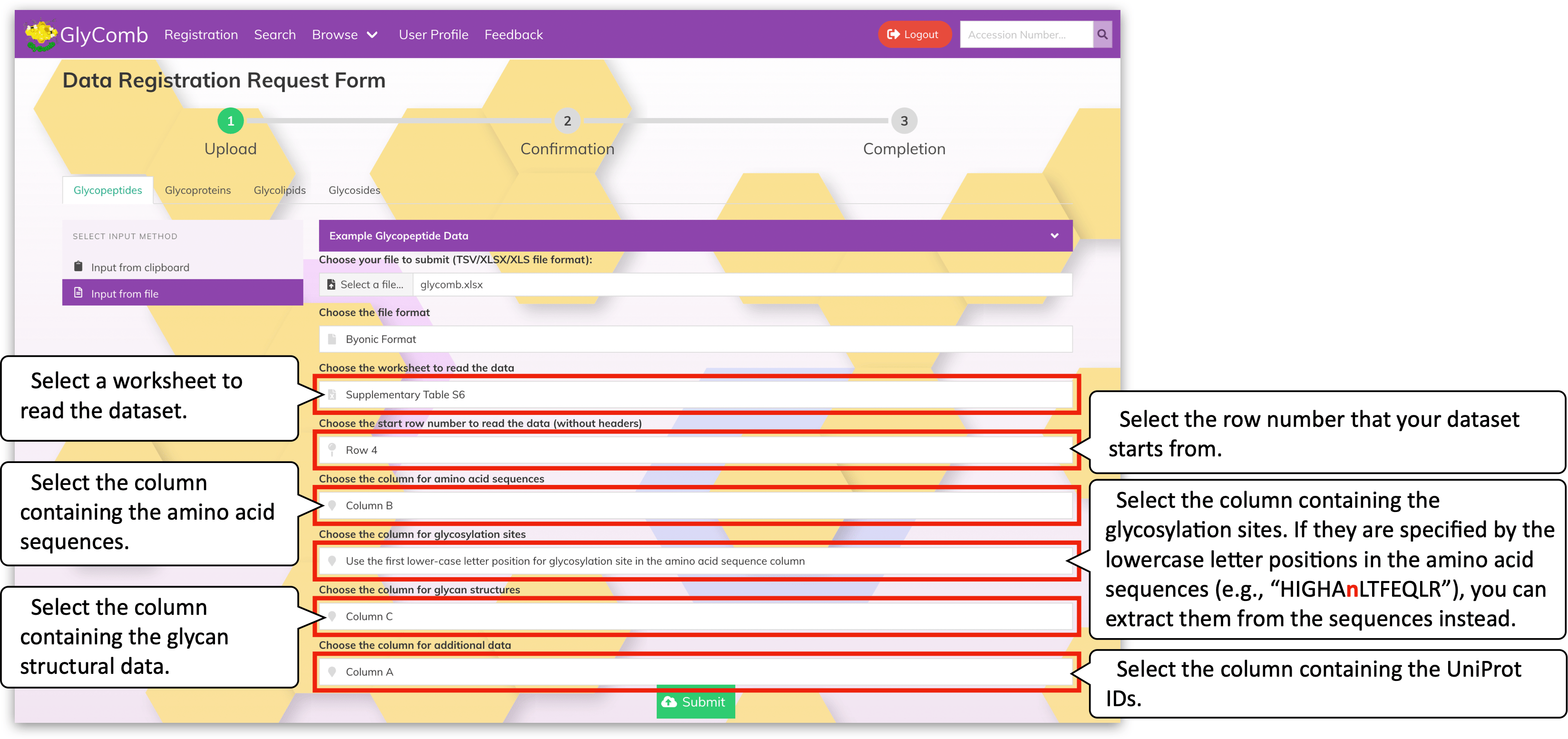
As you can see, the embedded glycosylation site information is extracted by selecting the "Use the first lower-case letter position for glycosylation site in the amino acid sequence column" item for the "Choose the column for glycosylation sites" field.
You can submit the configured dataset by clicking the green "Submit" button at the bottom of the screen. Then, the dataset validation result is listed on the next screen.
IMPORTANT: Your submission is not accepted to GlyComb yet.
In this screen, please verify that all of the status icons on the left-hand side show a green checkmark. There are possibly input issues if you see the red or orange icons. However, an orange icon shows a kind of warning, e.g., redundant input lines, not an error. Because GlyComb will automatically remove the redundant input lines, you can submit the dataset successfully in such cases. As a result, the same datasets will be assigned the same GlyComb accession number.
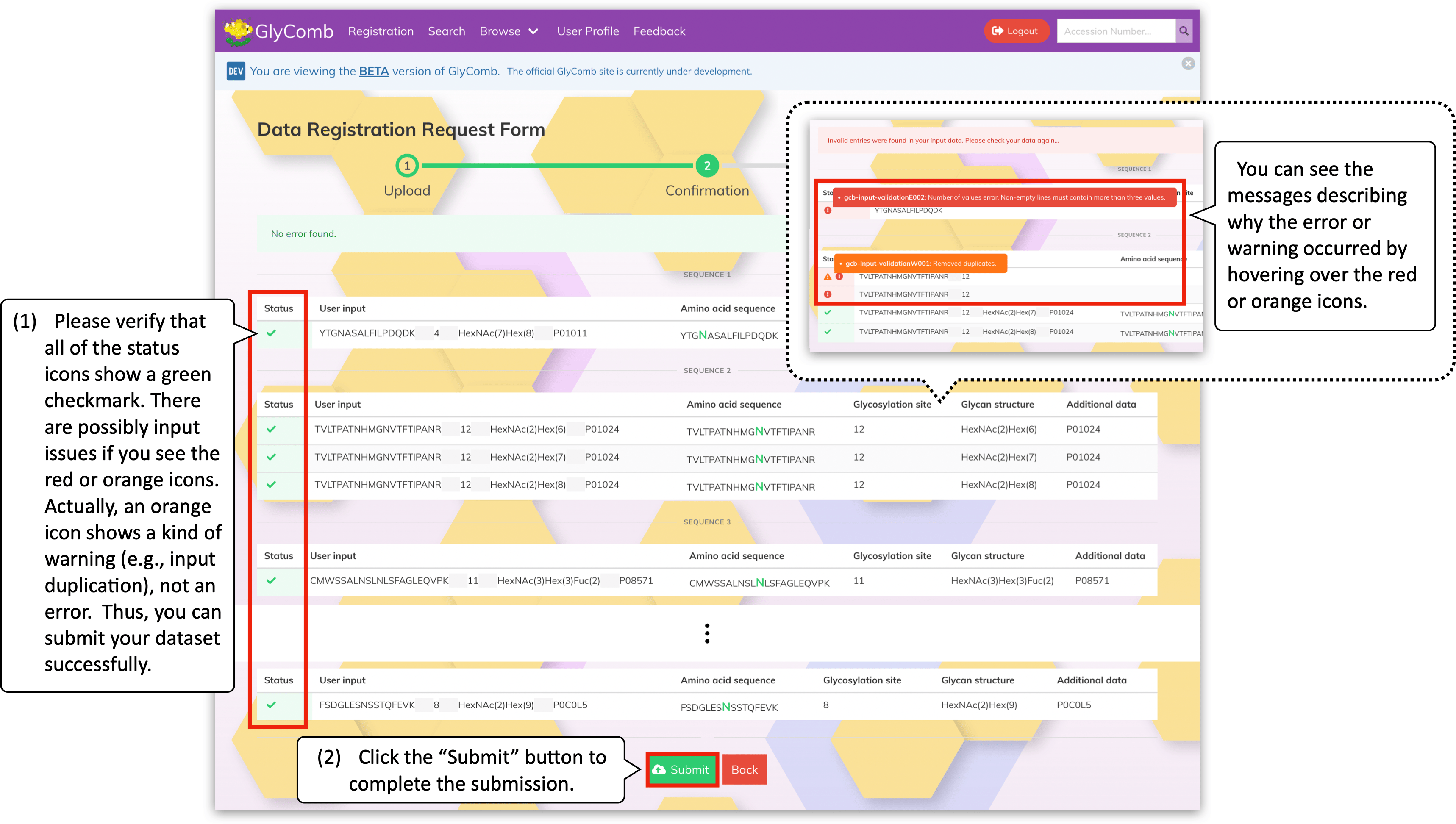
By clicking the "Submit" button at the bottom of the screen, GlyComb will accept your submission and the submission process will complete.
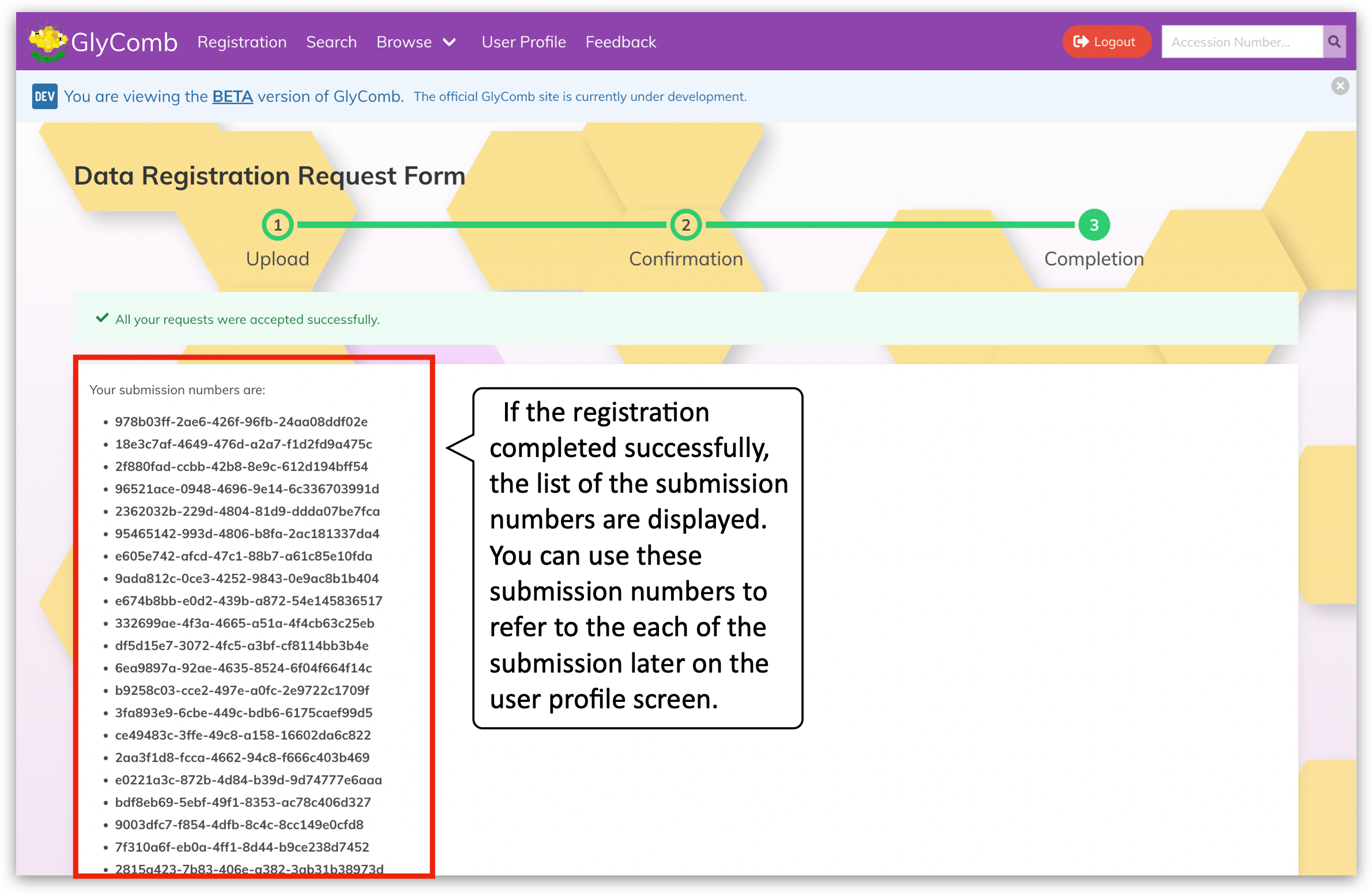
As a result of the submission, you will get the corresponding submission number. You can use this number to refer to each of the submissions later on the user profile screen.
¶ How to publish your submission
In GlyComb, your submission will not be opened to the public automatically.
Therefore, we want you to publish your datasets manually. GlyComb will assign the accession numbers to each submission in a batch process. Therefore, it will take several hours to assign an accession number to your submission.
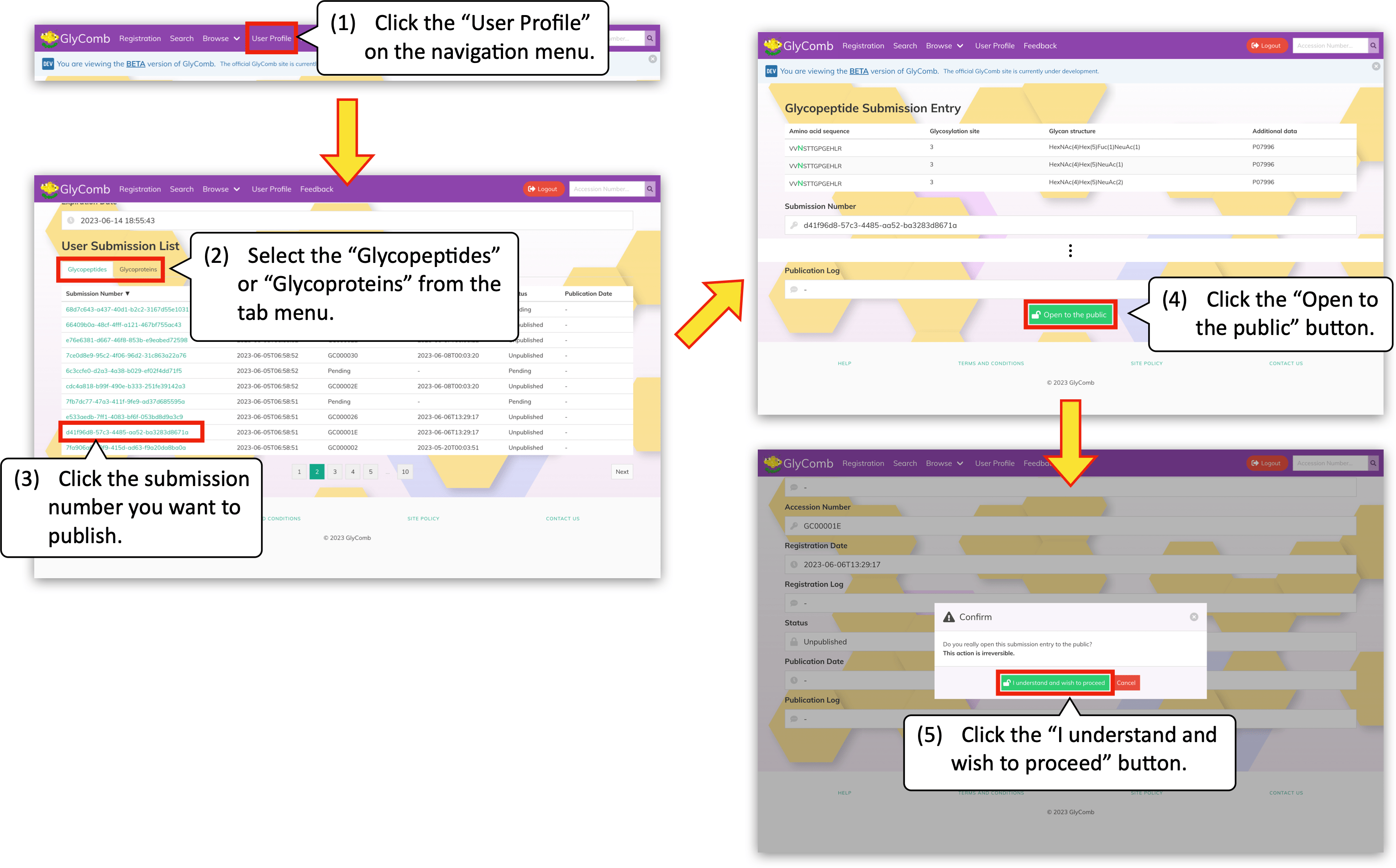
By clicking the "User Profile" on the navigation menu, you can browse your previous submissions. You can check whether an accession number is already assigned to your submission or not by looking at the "Accession Number" column in the submission list table on this screen. The accession number format in GlyComb is like GCxxxxxx.
If your submission is assigned to an accession number, please click the submission number in the list to publish the corresponding dataset. You will see the green "Open to the public" button at the bottom of the next screen. By clicking the button, a confirmation dialog will appear. You can publish the submission immediately by clicking the "I understand and wish to proceed" button in the dialog.
¶ Sending a feedback
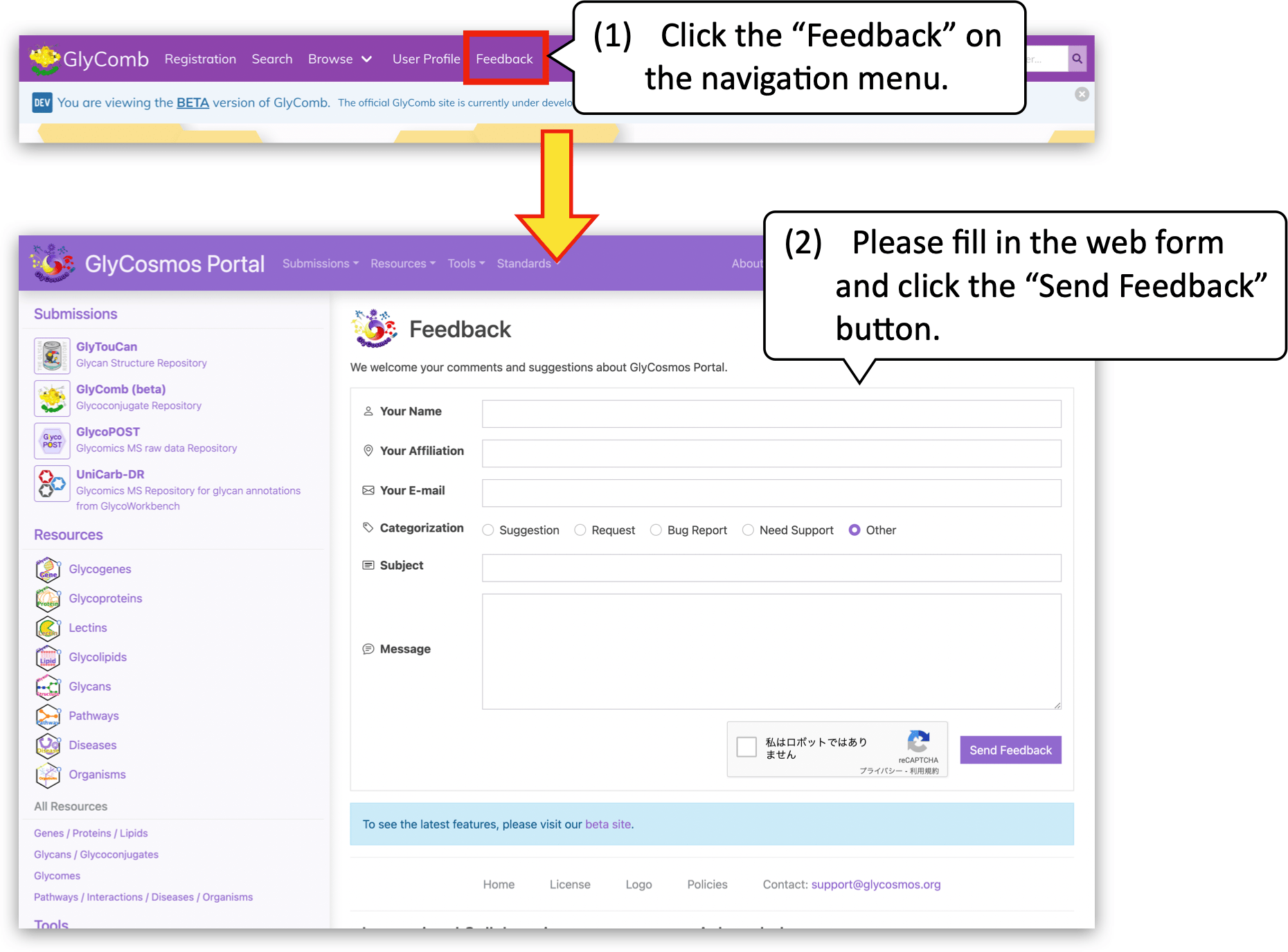
Your feedback and suggestions would be highly appreciated! Please feel free to send us your feedback from the web form on GlyCosmos portal.
Link to the feedback form: https://glycosmos.org/feedback
¶ References
- Perez-Riverol, Yasset, et al. "The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences." Nucleic acids research 50.D1 (2022): D543-D552.
- van Wijk, Klaas J., et al. "The Arabidopsis PeptideAtlas: Harnessing worldwide proteomics data to create a comprehensive community proteomics resource." The Plant Cell 33.11 (2021): 3421-3453.
- Fujita, Akihiro, et al. "The international glycan repository GlyTouCan version 3.0." Nucleic acids research 49.D1 (2021): D1529-D1533.
- Watanabe, Yu, et al. "GlycoPOST realizes FAIR principles for glycomics mass spectrometry data." Nucleic Acids Research 49.D1 (2021): D1523-D1528.
- Zhu, Jing, et al. "Quantitative longitudinal inventory of the N-glycoproteome of human milk from a single donor reveals the highly variable repertoire and dynamic site-specific changes." Journal of proteome research 19.5 (2020): 1941-1952.